Table of Contents
Authored by:
Abraham Nieva de la Hidalga 1, C. Richard A. Catlow1, Brian Matthews2
1UK Catalysis Hub, Research Complex at Harwell, Rutherford Appleton Laboratory, R92 Harwell Oxford Oxfordshire OX11 0FA
2Scientific Computing Department STFC, Rutherford Appleton Laboratory, Harwell Campus, Didcot, OX11 0QX, UK
Download a .pdf copy of this summary report
Download a .pdf copy of the full detail report
Key focus and activity
The main idea is to determine how easy it is currently to look for experimental data for reproducing experimental results, comparing different processing tools, and designing additional tools for processing and reproduction. The initial target area is X-Ray Absorption Spectroscopy (XAS) data processing. In this case, the publications of the UKCH were analysed to determine:
- which publications are linked to published data which could be used for reproducing results presented,
- what software is needed for reproducing the results
- what alternative software combinations could be used to obtain the same results
- which are the current barriers to effectively finding data, reproducing results, and further reusing the data in other experiments.
Area of Physical Sciences covered
The research projects of the UK Catalysis Hub are mainly aligned with the Catalysis [1] research area which to date has allocated more than 129 million pounds to 143 grants. In the current phase, the UK Catalysis Hub targets three research areas: (1) Optimising, Predicting and Designing New Catalysts, (2) Catalysis at the Water Energy Nexus, and (3) Catalysis for the Circular Economy and Sustainable Manufacturing.
Related PS research areas
Catalysis is a fundamental research area that has relevance to other areas including Analytical Science (spectroscopy and spectrometry techniques used), Carbon Capture and Storage (relevant projects impacting capture and reutilization of carbon derivates), Chemical Structure (analysis of the composition and use of catalyst in different areas), Combustion Engineering (catalytic converters, production of fuels from carbon and waste, biofuels), Energy Storage (fuel cells, reuse of waste products for fuel production), Materials Engineering – Composites (research of creation of catalysts and optimisation of their structure). Materials for energy applications (fundamental research on biofuels, use of waste materials, carbon capture and reuse for clean fuels).
Applicability to the Research Data Lifecycle
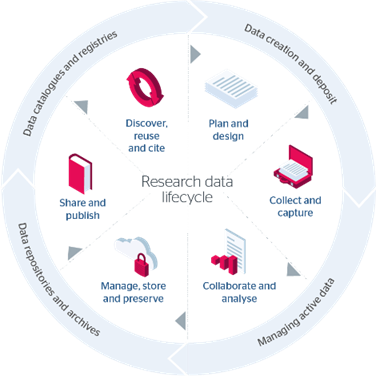
This case study offers a view of the support required for effectively preserving data so that it can be later discovered and reused. As such, it has direct links to the last three phases of the JISC research data lifecycle (Figure 1) [2]: (4) Manage, Store & Preserve, (5) Share & Publish, and (6) Discover, Reuse & Cite. However, the requirements derived from this case study will be relevant to all stages of the research data lifecycle.
Main outputs
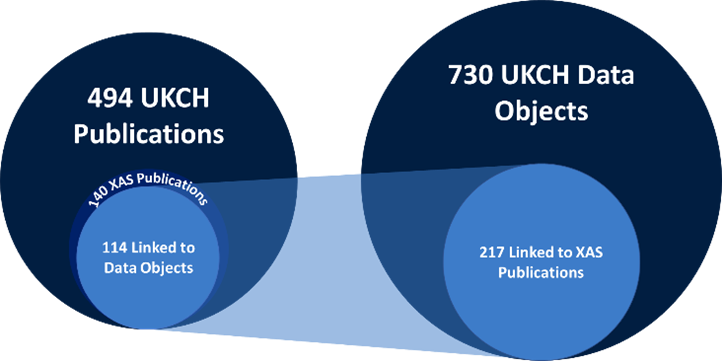
In this case study, we want to show how the combination of data from existing repositories can enable research reproducibility, data reuse, reuse of processing software, and development of advanced processing tools (combining workflows, machine learning and artificial intelligence). The initial target is XAS data processing because XAS analysis is a widely used technique for the characterisation of catalysts in ex-situ, in-situ and operando conditions. The relevance of XAS analysis for UKCH can be seen from the number of publications catalogued in the Catalysis Data Infrastructure (Figure 2). The indexing of data objects allows taking advantage of existing platforms which researchers use to preserve and publish catalysis research data such as STFC eData, ISIS TopCat, CCDC, and university research repositories. Therefore, the plan is to discover data which may support reproduction of results, paying particular attention to the types of data published, the software required for reproducing results presented and the processing and analysis tools which can be leveraged to facilitate these activities.

The CDI operates in a complex environment as a catalogue linking data objects to publications, providing context information which can enable reproduction, reuse, and extension of results. This case study will highlight what works, what can be improved and what are the pending tasks to better support these activities. The results of these will be directly relevant to the development of the PSDI as it will highlight some of the requirements for facilitating data discovery and reuse.
Outcomes and Recommendations
The initial goal was to analyse a set of publications and determine how well published data can support the reproduction, further use of results, and extension. Performing these types of activities with minimal human intervention (automated or semiautomated) is a common goal of many similar research infrastructures (e.g, NOMAD). However, the results obtained so far indicate that publishing and reuse of data requires significant overhaul and support, particularly requiring creation of tools that can simplify the management of data, including its publishing and referencing.
Outcomes
The types of data published are not always supportive of reproduction. The levels of published data vary considerably from additional tables and figures in document formats (supplementary data), to comprehensive data sets including raw data, intermediate results, and processed data. The exploration of the data indicates that most of the published data objects are supplementary data, while processable data is a small proportion of all published data.
| Table 1 – Data Objects linked to XAS publications | ||
| Type of Data | Count | % |
| All Data Objects indexed by CDI | 730 | 100.00% |
| Linked to XAS publications | 217 | 29.73% |
| Mention XAS data objects | 46 | 6.30% |
| Mention processable XAS data objects | 12 | 1.64% |
| Contain processable XAS data objects | 9 | 1.23% |
Moreover, when working with the processable data and trying to reproduce the experimental results, additional data was required in some instances and these data were not published or correctly referenced by the publications. Derived from these results compiled the following recommendations.
- Publish and link (reference) raw and intermediate data
- Publish metadata in structured form (XML, JSON) (in addition to document format), including:
- type of data published (.dat, .txt, .csv, .xlsx, .nxs, .opj, .opju, .pdf)
- software used to produce/read data
- link to additional data
- mapping of data to published results
- Prefer open source software (some data can not be processed without a licensed program e.g. Origin files (.opj/.opju))
These recommendations were discussed with UKCH stakeholders to try to discover the barriers to publishing research data. In this discussion, users highlighted three common issues:
- Problem 1: Difficult to use repositories. Usability was mentioned as a relevant issue. Users pointed out that the processes and tools which support data publication are hard to use and not intuitive, requiring users to know how to annotate, format, and curate the data.
- Problem 2: Keeping track of processing tools and intermediate results. The tasks of mapping published results is not well supported. From collection at the lab or experimental facilities, through processing and analysis, to finally formatting and publishing keeping track of provenance is a manual process. There are no current tools supporting it.
- Problem 3: Lack of guidance on which data to publish. Entities requiring publishing of data do not specify the kind of data to be published, consequently there is a wide range of data object types being published. Some authors are very thorough and publish raw, intermediate, and processing data and map it to published results; meanwhile, other authors just include additional (supplementary) data in the form of documents and figures.
The following requirements are intended to address these problems.
- Tools that generate provenance metadata to trace the processing trail for each result presented. This could be a modification of existing logging features, aligning them with standards like PROV-O.
- Processing and analysis tools which produce outputs ready for publishing (with metadata and provenance data ready). Experiment management systems already encode metadata about experiments. Processing and analysis tools could take provenance and setup data to produce new links in the provenance chain and additionally produce metadata for publishing
- Simplify the deposition of data, include the process in existing workflows, transparent to users. The tools outlined above, could produce outputs which if fed to deposition systems could simplify data curations and publishing tasks.
- Data annotation tools which can produce metadata for mapping results to source objects. These tools should be integrated into the data publishing facilities to support the publication of data with prefilled formats and recommendations to support deposition.
During the discussion, the following additional concerns were raised by users.
- Problem 4: Data Ownership. Issues about the ownership of data were also raised, this is related to clear data management policies and use agreements. Most times this has been addressed by licencing agreements.
- Problem 5 Entry of experimental setup data and metadata in physical lab books. Facilities and laboratories still rely on manual input of information on lab notebooks. Information in those is harder to retrieve.
- Problem 6 Lack of programmatic (API, web services) access to repositories. Repositories provide mainly manual interface for accessing data. This prevents use of high throughput methods.
Addressing these additional issues requires further analysis and discussion with stakeholders. These could be further covered in the PSDI design phase.
Full detail report
The full detail reported produced by this case study can be found at: Full Report Link