‘We are laying the foundation for the next generation of AI-drive scientific discovery.’
Dr James Gebbie-Rayet and his research team are poised to revolutionize the Biomolecular Simulation Research field by addressing long-standing, systemic challenges. James explains, ‘We often don’t know what simulations have been done before. If we had a database of simulations, we’d stop duplicating work and start building on what already exists.’ Through developing BioSimDB – included in PSDI resources, James and his team aim to overcome these challenges.
The challenge – Transforming Biomolecular Simulation’s Research Culture
‘The most important bit, in my opinion, that goes missing is how that simulation was created, because that’s the bit that matters the most.’
Scientific research thrives on reproducibility. Yet, within Biomolecular Simulation, a fundamental flaw persists: a lack of clear, standardized methods and the infrastructure to document and share the simulations intricate steps.
Traditionally, researchers publish their findings based on final simulation outcomes without detailing the precise methodologies and parameters used to achieve those results. ‘People in academia are under pressure to publish quickly, that’s why they don’t prioritize sharing their full methodologies. We need to change that mindset.’ This missing information makes it nearly impossible to replicate studies, verify discoveries, or build upon prior simulations.

Inspiration
This became all too evident for James, during the COVID pandemic, ‘we established a task force, under HECBioSim, sharing resources nationally [to support drug discovery]. We quickly realized that we couldn’t do it properly because some components required were missing in our field.’ This experience inspired James and the CCPBioSim to take matters into their own hands. ‘The pandemic made us realize that if we don’t fix the reproducibility problem now, the next crisis will hit, and we’ll be stuck in the same situation – scrambling to understand each other’s research with missing information.”
His aim was to create data provenance tools that, once installed, automatically capture and archive every step of the simulation process, in a readily understandable and shareable format.
‘COVID really exposed the cracks in our field. We needed to share simulations across different groups, but there was no infrastructure to do it efficiently. That’s why this resource is so important now.’
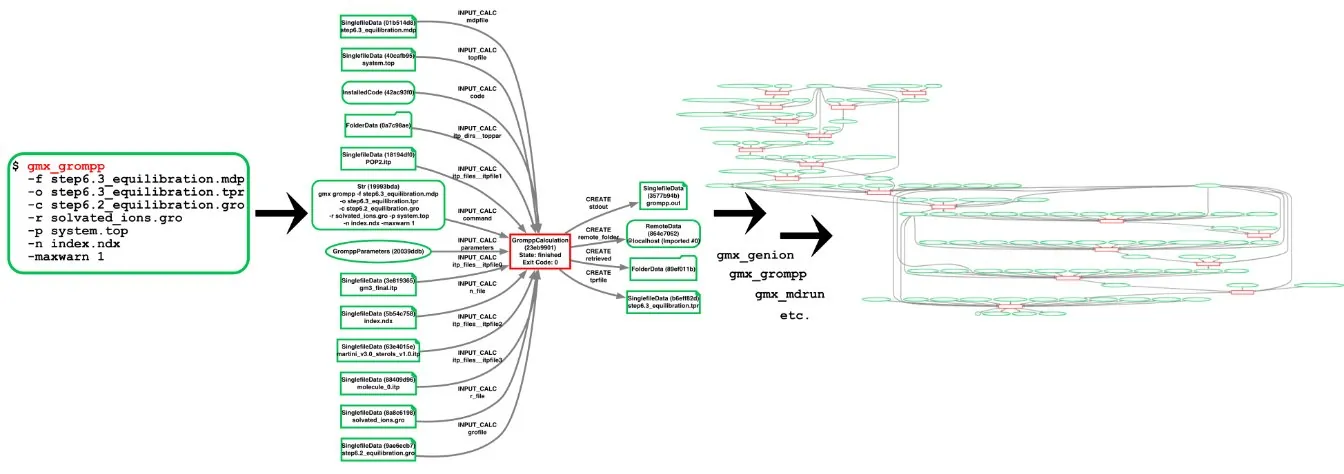
Input and output node network produced when performing simulation steps via aiida-gromacs.
This resource not only ensures that researchers share their results but the vital, underlying processes that led to them, fostering a more robust and accountable research culture. ‘Ultimately, this is about making scientific research more sustainable. If we don’t start tracking and sharing our data properly, we’re just reinventing the wheel over, and over again.’
The solution – Built with researchers in mind
One of BioSimDB’s greatest strengths is its ability to seamlessly integrate with existing workflows. Researchers no longer need to manually document their methodologies or input data into separate repositories – the BioSimDB does this in the background, automating data capture and standardizing results. ‘Researchers don’t have to change how they work – but they get all the benefits.’ Reducing the administrative burden and allowing scientists to focus on the actual science rather than documentation.
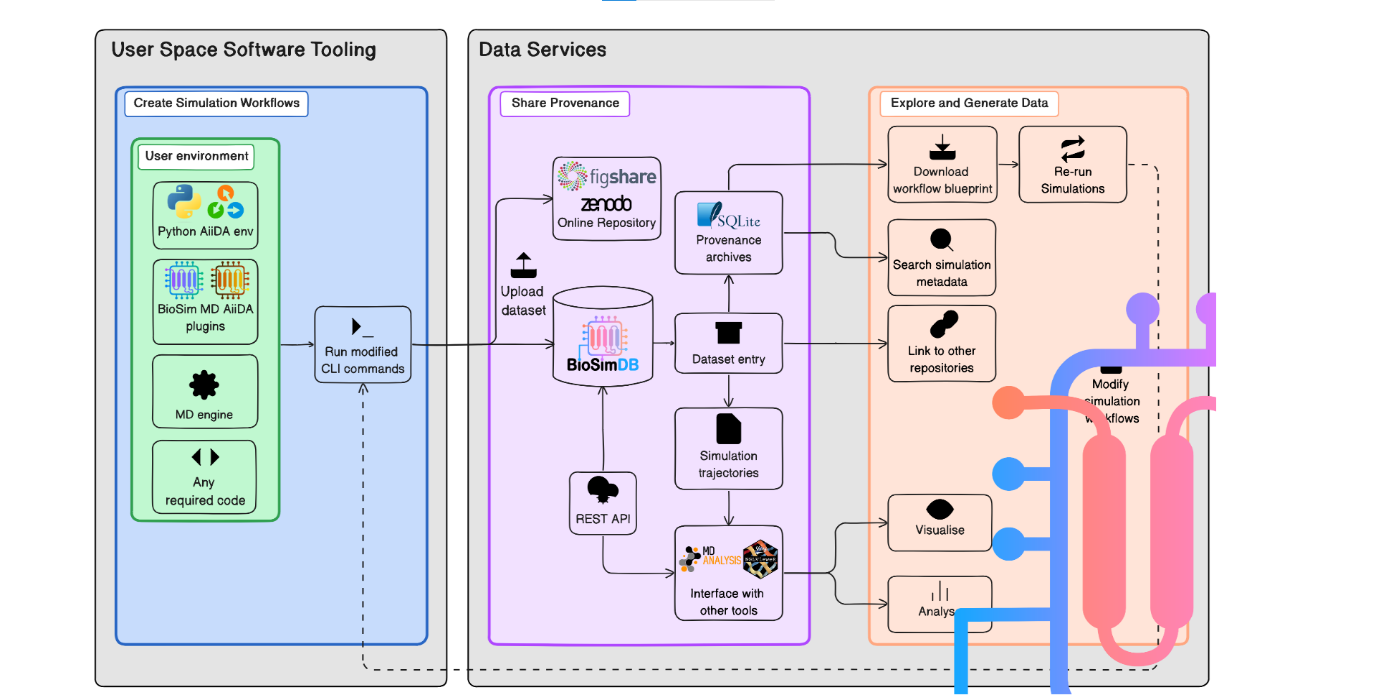
Vision to produce and share biomolecular simulation workflows and data.
‘We design these really complicated simulations, but the physics is actually very simple, the complexity comes in the process, and that’s what we need to track.’
The full provenance of a simulation is then accessible – from experimental structures and force field configurations to computational parameters – everything is meticulously recorded. Research findings become, not only verifiable but reusable in ways previously impossible. This level of efficiency benefits not only individual researchers but also institutions and funding bodies by optimizing research outputs.
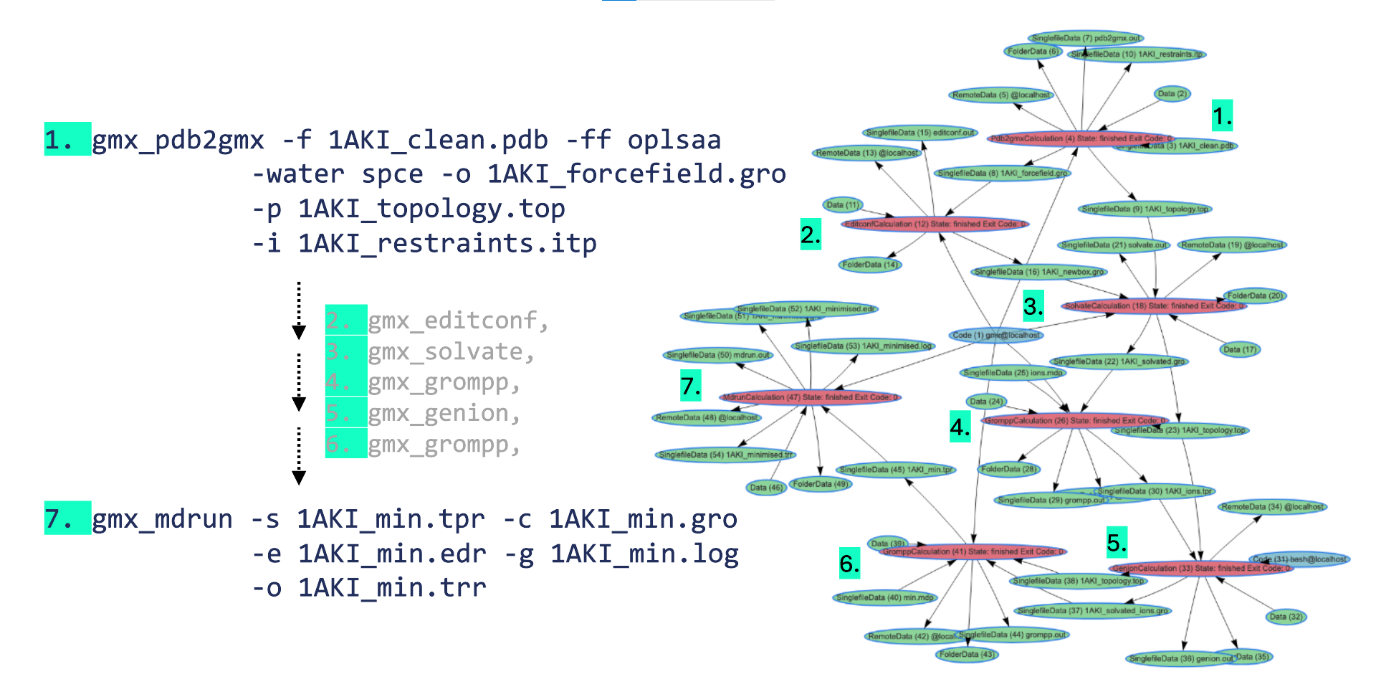
Example of using modified gmx commands via aiida-gromacs (left) to build a provenance graph (right)
Furthermore, ‘We’re building a database repository so we’ll have this huge resource of data that other people have run, which means we can stop duplication of effort.’ This repository will be accessible, enabling researchers to save valuable time, computational resources, and will democratise simulation informed research.
Through eliminating the opacity that often clouds Biomolecular Simulation research, BioSimDB positions itself as a transformative force.
Global Impact
‘Right now, research groups around the world are duplicating work without realizing it. If we can centralize these efforts through this, we can make real progress in tackling global health challenges.’
By enhancing the reliability of Biomolecular Simulations BioSimDB has the potential to accelerate breakthroughs in drug discovery, ‘The ability to accurately reproduce biomolecular simulations means we can accelerate drug discovery – reducing the time it takes to go from a theoretical compound to a viable treatment… every step in a simulation is recorded, meaning pharmaceutical companies and researchers can trust the results they’re working with when developing new medicines… If we can validate our simulation models with high confidence, we can reduce reliance on expensive and time-consuming physical trials, ultimately accelerating medical innovation.’ insilicoUK is a political movement working on the policy elements of this development.
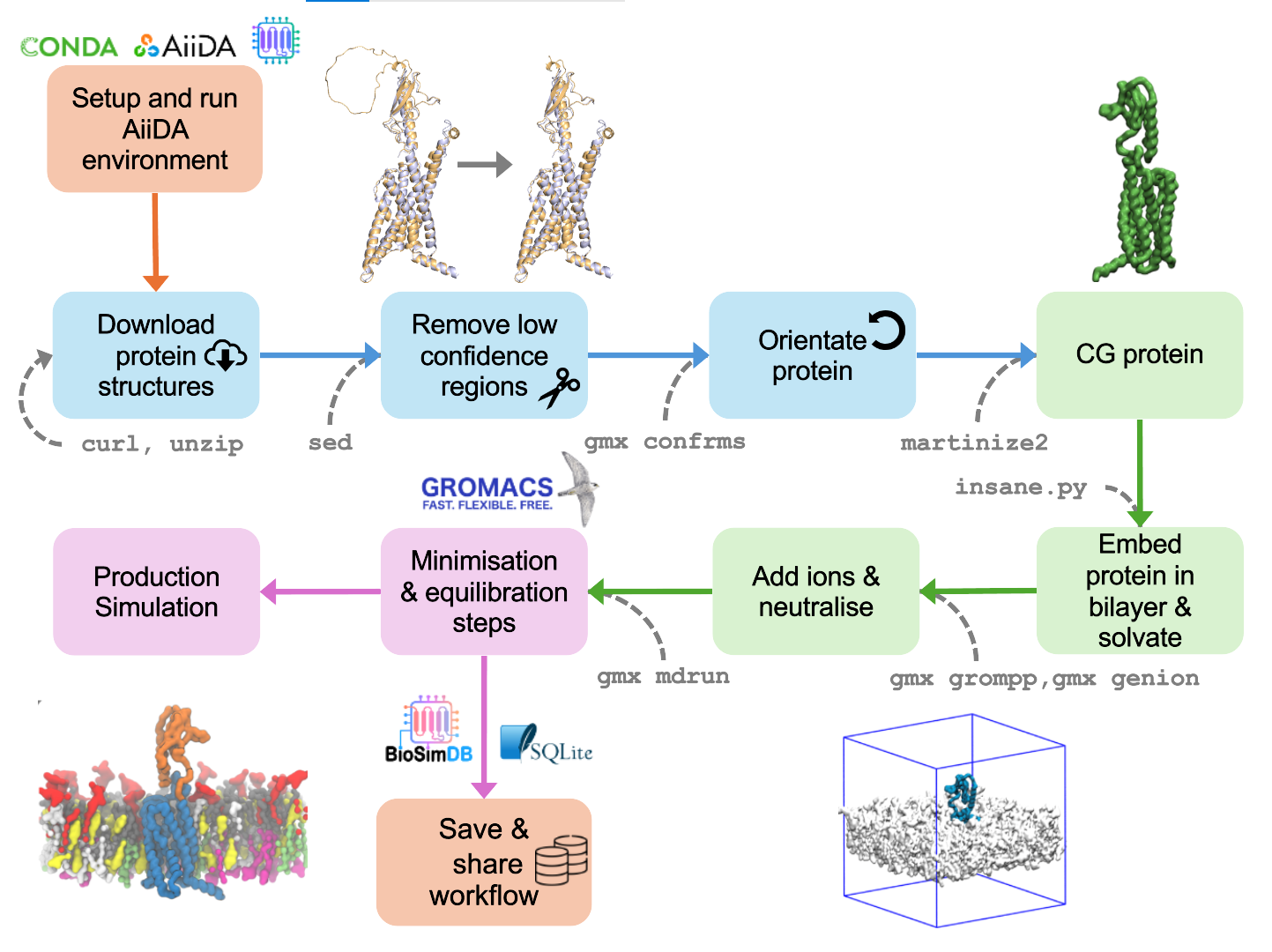
Caption: Schematic example of the multiple simulation setup of a coarse-grained protein embedded in a membrane, captured with aiida-gromacs
‘We talk about drug discovery, but this is bigger than that.’
Disease research will also be positively impacted. ‘It will allow us to model diseases, study protein malfunctions, and understand the molecular basis of conditions in a way we’ve never been able to before, that is based on trust and accountability… Being able to track every detail of a biomolecular simulation means we can improve processes around personalized medicine, tailoring treatments at the molecular level with greater accountability.” The ability to reproduce and validate simulations with high confidence means that AI-driven drug design, molecular modelling, and biomedical research can advance at unprecedented speeds.
In a world where computational power is increasingly shaping scientific progress, ‘AI in drug design is limited by poor-quality datasets. We’re creating a gold standard in simulation data that AI can train on, making predictive modelling more reliable… By ensuring that data is standardized and high-quality, we are laying the foundation for the next generation of AI-driven scientific discovery.’ ensuring that this power is harnessed with integrity, efficiency, and transparency.
Conclusion
As part of PSDI, the BioSimDB for Biomolecular Simulation significantly enhances the organisation’s offering by tackling data integrity and reproducibility. While PSDI focuses on data sharing and collaboration, this resource expands PSDI’s mission by introducing a structured, scalable approach to simulation data management. Aligning with global movements toward open science and FAIR data principles. Making Biomolecular Simulations more sustainable, credible, and collaborative.
‘The technologies we are building now, will shape the next decade of biomolecular research, ensuring that findings are credible, reproducible, and impactful.’
With its promise to reshape research culture, and drive innovation, the BioSimDB for Biomolecular Simulation – PSDI resource, is more than just a resource, it is a movement towards a more open, accountable, and impactful scientific future.
Credits:
Jas Kalayan, responsible for the development of BioSimBD tools. Gemma Poulter’s team, and particularly Andrew Harper, who have developed the data repository, based on InvenioRDM.